RESEARCH
I investigate brain function with modelling and analysis methods, ranging from the simulation of intracellular physiology to analyses of whole-brain dynamics. The main focus of my research is the neural basis of cognition and consciousness, with a view to understanding these phenomena in health and disease, and to identifying the computations required to support them in machinery. In this regard, I believe we're a long way from developing artificial general intelligence and I'm interested in the qualitative leaps required to get there. I work closely with experimentalists using a wide range of techniques, including in vitro and behavioural electrophysiology, behavioural psychology and neuromaging methods (fMRI, EEG, MEG). I record data myself from time to time.
Perspective
Computational models explain and predict data. If they don't do these things, they can reasonably be described as useless for brain science - though potentially useful for engineering. The late David Marr referred to three levels of analysis for modelling brain function, but I believe two levels are sufficient - abstract models characterise the computations underlying brain function and neural models address the implementation of those computations in brain circuitry. Analytic methods can be used to relate these models to one another, enabling us to investigate the relationships between data recorded with different methods, from different subject groups, and from different species. This approach assumes that neural data is mutually informative across scales and highlights the need for re-analysis of openly available data. Money is scare in science (and elsewhere, of course) so this kind of efficiency is crucial. Ultimately, understanding neural computation requires an understanding of the roles played by intracellular physiology at the systems level. Thus, this perspective emphasises modelling and analysis across scales.
Brain circuit modelling
|
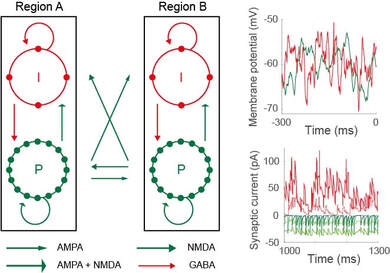
I have a lot of experience with biophysically-based network models, that is, network models whose variables and parameters have direct anatomical and physiological correlates. In particular, I've developed many models with cellular and synaptic resolution, allowing me to investigate the computations revealed by single-cell recordings from behaving animals, and those revealed by local field potentials (LFP) and electroencephalogram (EEG) data (these signals can be approximated from simulated synaptic currents). The upshot - if a model can simultaneously account for neural and behavioural data, then it offers an explanation of how neural circuitry supports the behaviour. The key is to make testable predictions for. The figure shows a model schematic (pyramidal neuron P, inhibitory interneuron I) alongside simulated cellular and synaptic recordings (adapted from Standage et al, J Vis, 2019)
|
Future directions Biophysically-based network models have tremendous scope for clinical work, enabling the simulation of dysfunctional physiology, such as the degradation of cellular morphology (e.g. aging), the abnormal expression of neurotransmitter and neuromodulator receptors (e.g. schizophrenia), the depletion of neuromodulatory neurons (e.g. Parkinson's disease) or imbalances of excitation and inhibition (e.g. epilepsy). They further enable the simulation of pharmacological intervention in the treatment of neuropathologies, and of behavioural experiments with patient populations. This approach is often referred to as computational psychiatry. If you're a clinician and these things are of interest, please contact me.
Network neuroscience
|
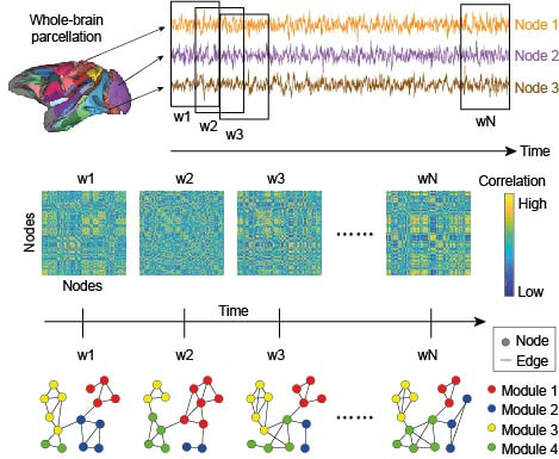
Anything that can be described as a set of entities and relationships can be represented as a graph. The entities are nodes and the relationships between them are edges or connections. In recent years, the use of graphical methods to construct formal networks from fMRI signals has provided a platform to investigate neural information processing over a broad range of spatial scales, including whole-brain computation. Under this approach, brain regions are nodes and correlations in their activity constitute edges, the assumption being that if two regions are co-active, then they're working together as part of a functional network. There are many ways to use these methods, and I frequently use them to quantify the time-evolution of functional networks during cognitive tasks. Associations between the statistics of network properties and task performance provide clues about the whole-brain bases of cognition. In particular, I'm interested in whole-brain modular composition, i.e. how the sub-systems of the brain can be mixed and matched to support the task at hand. There is growing evidence that we do exactly that, but the mechanisms by which we do it are poorly understood. The figure provides an overview of this general approach (adapted from Standage et al, Cereb Cortex, 2020)
|
Future directions I want to use this approach with EEG and MEG, to investigate whole-brain computation in real time. EEG and MEG have millisecond resolution, so it's doable with enough computing power (it can be described as an embarrassingly parallel problem). If you're interested in collaborating in this area, please contact me.
Executive Control
|
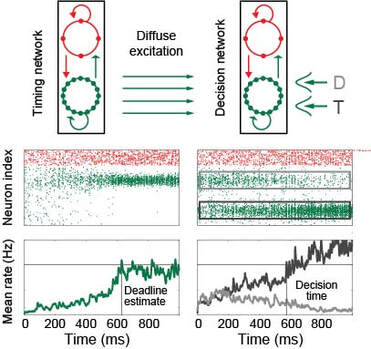
From my perspective, executive control refers to the ability of one brain circuit to influence the computations performed by another. Much of my research investigates the dynamics of modulatory signals and their target circuits, e.g. the mechanisms by which we encode elapsed time and how these signals influence decision making. More recently, I've begun to focus on distributed brain circuitry, e.g. the ability of brain regions (or groups of regions) to drive changes in whole-brain network states. I'm particularly interested in the source of executive signals, such as the emergence of structured neural activity in the absence of sensory stimuli, and the computations supported by layer-specific connectivity in hierarchical circuitry. In the figure, a neural estimate of elapsed time controls speed-accuracy trade-off by modulating the dynamics of decision circuitry. T target, D distractor (adapted from Standage et al, PLOS Comput Biol, 2013)
|
Decision Making
|
Decision making refers to the deliberation over evidence for the purpose of making a choice. A lot of data suggest that decisions are made by accumulating evidence until a running total (a decision variable) reaches a threshold, the crossing of which drives choice behaviour. There is ongoing debate about the nature of such bounded accumulation, e.g. how much the decision variable decays while it accumulates. In much of this work, different groups have characterised the computations underlying behaviour with different models, the principles of which have been proposed to support decision making. The problem here (in totality) is that decision making isn't monolithic, so that each group may be correct that its model captures the relevant principles in the context of its experiments. In my opinion, debates about Model A vs Model B are less interesting than the mechanisms by which we have flexible control over decision circuitry, allowing us to be A-like or B-like in different contexts. Thus, my research in this area is founded on the premise that we have control over neural dynamics on a range of spatial scales (local circuitry, systems of local circuits, whole-brain distributed processing). As such, our circuitry can be leaky when accumulating evidence would be an impediment to choice behaviour (such as in rapidly-changing environments), can be impulsive (amplifying a decision variable) when any choice is better than none, and can be something like perfect when we have the time and focus to discriminate carefully between alternatives with widely varying outcomes. In most of this work, I investigate the ways that modulatory signals control processing in different contexts.
|
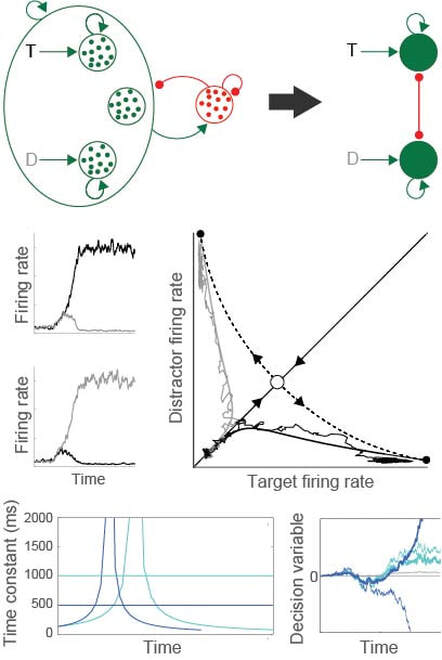
The figure depicts the reduction of a biophysically-based network model to a simplified, 2-variable system, tractable for formal analysis (middle row). Under this approach, our models make context-dependent (dark and light blue) predictions for decision tasks, based on the properties of circuit dynamics (adapted from Standage et al, Front Comput Neurosci, 2011; Standage et al, Front Neurosci, 2014)
Learning and Memory
|
Learning and memory refer to many overlapping phenomena. For my purposes, learning refers to the formation of memory, and memory refers to the encoding of associations between things, so they can be recalled (e.g. facts) or executed (e.g. motor skills). These definitions are purposefully general, encompassing neural processes at the intracellular and whole-brain levels and their support of adaptive behaviour. At the intracellular level, I'm interested in biophysical processes that quantify correlations between pre- and post-synaptic neural activity, and the mechanisms by which these physiological variables influence the strength of synapses (synaptic plasticity). At the whole-brain level, I'm interested in how adaptive behaviour is supported by the reconfiguration of functional networks.
Future work I'm interested in how synaptic plasticity supports changes to whole-brain functional networks during learning, i.e. integrating the above two approaches. Thus, I'm especially interested in the use of reinforcement learning with network models grounded in anatomical principles. If you're an RL person and that sounds interesting, please contact me. |
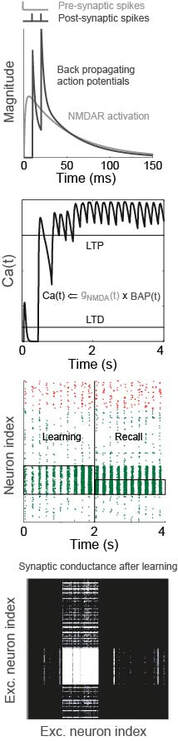
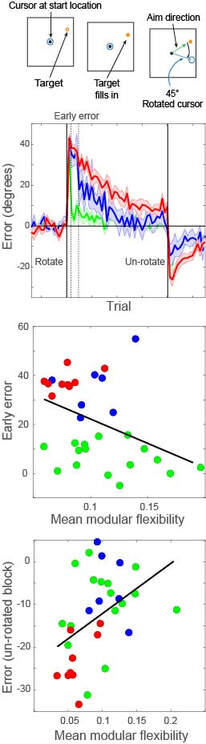
In the modelling figure, a calcium-like variable captures pre- and post-synaptic spiking correlations, via the interplay between NMDA receptor activation and back-propagating action potentials (BAP). This physiological model of plasticity is embedded in a model of hippocampal region CA3, which learns to complete an "index" pattern during theta-band oscillations (adapted from Standage et al, PLOS ONE, 2014). in the behavioural figure, three groups of participants learn a visuomotor rotation task. Participants whose whole-brain networks show more flexible modular composition are faster learners (Standage et al, in prep)
Working Memory
Working memory refers to the active retention and manipulation of information for several seconds. Given the fundamental importance of working memory to cognition, its storage limitations are surprisingly severe. My research in this area addresses the neural basis of these limitations and the strategies used to mitigate against them. Behaviourally, storage limitations are typically measured in terms of the number and precision of memoranda, i.e. how many things we can keep in mind and how precisely we can keep them there. There is fierce debate about whether these measurements reveal limitations on the number or resolution of encoded items, but our neural simulations predict that both measurements are manifestations of the same underlying mechanism (competitive neural dynamics). My perspective on this issue is similar in principle to my perspective on decision making above, i.e. debates about Model A vs Model B are less interesting than the mechanisms by which we can be A-like or B-like in different contexts.
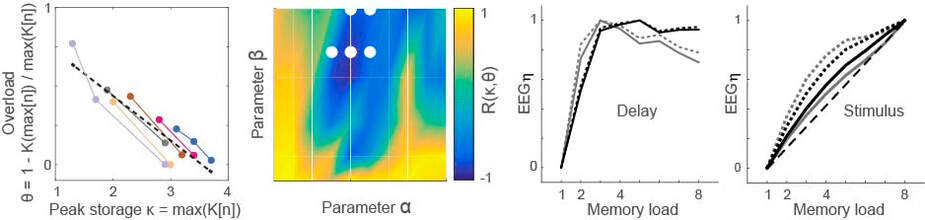
Working memory overload provides a window on strategies for storage. In the figure, overload is defined by task performance as a function of memory load (n). By searching over parameter values in a fronto-parietal model, we identified configurations that account for behavioural data on overload (blue) and other performance statistics (white dots). Under the identified configurations, the model accounts for EEG recordings during a memory delay and makes predictions for EEG signals during stimulus presentation (adapted from Standage et al, J Vis, 2019)